A couple of months ago I posted an overview of simple estimation of hierarchical events using python and petersburg. At the time it probably seemed a little bit trivial, just building a structured frequency model and drawing samples from it. But I have finally implemented the next step to complete the intended functionality. This post is a quick overview of how to use the new functionality, with more to follow on exactly how it works under the hood.
If you read through a few older posts on petersburg, you’ll recall that we represent the structure of a problem as a directed acyclic graph (with a few extra concepts added), where edges are weighted according to their likelihood to be traversed and have costs, nodes have payoffs, and we have some single ‘starting node’. This is all a bit abstract to think about, so let’s take a simple example: the weather.
Setting up the problem
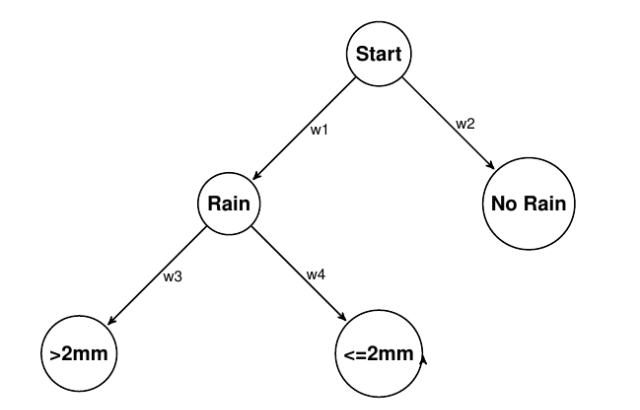
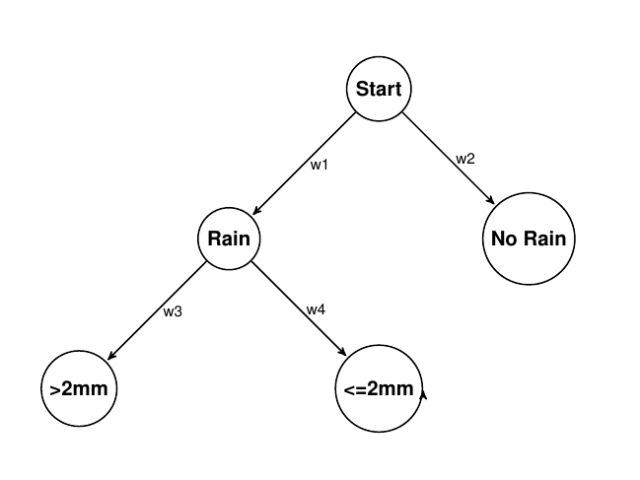
On any given day, it can rain, or not rain. If it does rain, then it can rain less than 2mm or more. So immediately, we’ve formed a simple little hierarchy. For the sake of estimation, we don’t really care that much about costs and payoffs as much as weights (at this point), so we will assume those are all 0. We can represent it as a graph like so:
The estimation task is to take some observation data and learn the appropriate weights {w1, w2, w3, and w4}.
Generating sample data
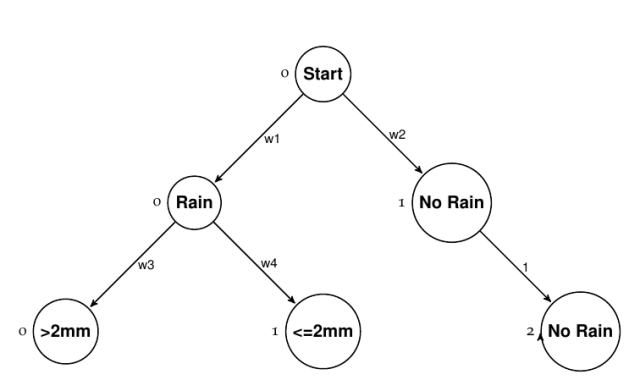
As was shown in the previous post, we can represent an observation in a tabular form by considering the graph to be layered. Since the path to each endpoint is not of equal length, we add a trivial node to even things out:
Notice that we also gave each node an ID within each layer. So we have three layers that together fully represent the problem. Each outcome is represented by:
- >2mm = [0,0,0]
- <=2mm = [0,0,1]
- No Rain = [0, 1, 2]
Using the simple frequency estimation, we don’t really need any feature data, but for now, let’s assume that we have some extra data along with each observation in the form of a feature vector. We can generate sample datasets pretty simply with some python:
def make_data(n_samples=10000):
_chance_of_percip = 0.2
_chance_of_heavy = 0.4
y = []
X = []
for _ in range(n_samples):
if random.random() < _chance_of_percip:
if random.random() < _chance_of_heavy:
y.append([0, 0, 0])
X.append([random.random() + 10 for _ in range(20)] + [random.random() * 0.2 + 1.1])
else:
y.append([0, 0, 1])
X.append([random.random() + 25 for _ in range(20)] + [random.random() * 0.2 + 2.1])
else:
y.append([0, 1, 2])
X.append([random.random() - 10 for _ in range(20)] + [random.random() * 0.2 - 2.0])
return np.array(X), np.array(y)
Notice that the feature vectors (X), have some values that do actually inform us about the outcome (y). That will come in handy later.
Frequency Based Estimation
Under the hood, there have been some changes made to the frequency estimator in petersburg, but it still functions the same. Using a scikit-learn style interface, you pass in some observations, the count of observations across each edge is used as it’s weight.
The prediction used to be one single simulation’s outcome, but now you can pass in num_simulations (which defaults to 10), and the most common outcome out of that batch of simulations will be returned.
Using petersburg’s FrequencyEstimator, we can train a model on some data from the above function, and validate it on a hold out set of the same data to see how well it works:
def validate(y, categories, truth):
t = 0
f = 0
y = y.reshape(-1, ).tolist()
final_idx = len(truth[0]) - 1
for idx in range(len(truth)):
truth_tuple = (final_idx, truth[idx][-1])
truth_idx = [k for k in categories.keys() if categories[k] == truth_tuple][0]
if y[idx] == truth_idx:
t += 1
else:
f += 1
return t / (t + f)
if __name__ == '__main__':
# train a frequency estimator
X, y = make_data(n_samples=100000)
clf = FrequencyEstimator(verbose=True, num_simulations=10)
clf.fit(X, y)
X_test, y_test = make_data(n_samples=10000)
y_hat = clf.predict(X_test)
# print out what we've learned from it
print('nCategory Labels')
labels = clf._cateogry_labels
print(labels)
print('nUnique Predicted Outcomes')
outcomes = sorted([str(labels[int(x)]) for x in set(y_hat.reshape(-1, ).tolist())])
print(outcomes)
print('nHistogram')
histogram = dict(zip(outcomes, [float(x) for x in np.histogram(y_hat, bins=[2.5, 3.5, 4.5, 5.5])[0]]))
print(json.dumps(histogram, sort_keys=True, indent=4))
accuracy = validate(y_hat, labels, y_test)
print('nOverall Accuracy')
print('%9.5f%%' % (accuracy * 100.0, ))
Which will give you the output:
Category Labels
{0: (0, 0), 1: (1, 0), 2: (1, 1), 3: (2, 0), 4: (2, 1), 5: (2, 2)}
Unique Predicted Outcomes
['(2, 0)', '(2, 1)', '(2, 2)']
Histogram
{
"(2, 0)": 8.0,
"(2, 1)": 57.0,
"(2, 2)": 9935.0
}
Overall Accuracy
79.86000%
The outcomes are of the form (layer, id), so you can see that in the 10,000 predictions made, 9,935 were made for ‘no rain’, with an overall accuracy of 79.86%. By changing the num_simulations to 1, the overall accuracy drops to 66.21%. (we will explore this parameter more in a separate post).
This model works decently, but we are totally ignoring that useful feature vector by only using frequencies.
Classification Based Estimation
Within a layer, the weights of each edge are related to the probability of that edge being traversed by a given agent (i.e., in a particular instance, which path is most likely to be taken). In the context of the whole graph, a single edge is a conditional probability. In that way, w3 is related to the probability of there being >2mm of rain given there being rain at all.
So it would be more clever to use an actual classifier to determine these probabilities using the feature vectors for each sample for each edge. We can consider w3 then to be a function of some vector, X. Petersburg now has an object that handles this for us, with the same interface as the FrequencyEstimator. The object is called MixedModeEstimation, but for this example, we will ignore the ‘mixed’ part of is, because every single weight in the problem will be modeled with a separate classier, a Logistic Regression model.
Everything else left equal, we can try out this method very similarly:
if __name__ == '__main__':
# train a frequency estimator
X, y = make_data(n_samples=100000)
clf = MixedModeEstimator(verbose=True)
clf.fit(X, y)
X_test, y_test = make_data(n_samples=10000)
y_hat = clf.predict(X_test)
# print out what we've learned from it
print('nCategory Labels')
labels = clf._cateogry_labels
print(labels)
print('nUnique Predicted Outcomes')
outcomes = sorted([str(labels[int(x)]) for x in set(y_hat.reshape(-1, ).tolist())])
print(outcomes)
print('nHistogram')
histogram = dict(zip(outcomes, [float(x) for x in np.histogram(y_hat, bins=[2.5, 3.5, 4.5, 5.5])[0]]))
print(json.dumps(histogram, sort_keys=True, indent=4))
accuracy = validate(y_hat, labels, y_test)
print('nOverall Accuracy')
print('%9.5f%%' % (accuracy * 100.0, ))
Which as you might expect, gives perfect results (because the feature vector has a trivial indicator in it):
Category Labels
{0: (0, 0), 1: (1, 0), 2: (1, 1), 3: (2, 0), 4: (2, 1), 5: (2, 2)}
Unique Predicted Outcomes
['(2, 0)', '(2, 1)', '(2, 2)']
Histogram
{
"(2, 0)": 7992.0,
"(2, 1)": 790.0,
"(2, 2)": 1218.0
}
Overall Accuracy
100.00000%
A note on mixed mode
In a future post I will go into the internals of the mixed mode estimator a bit more, but for now consider the case where an edge of the graph is traversed very rarely. Because each edge of the graph has it’s own discrete classifier, trained on all of the data that has gone through it’s parent node, some graphs with unbalanced traversals end up with edges that have insufficient data to model.
In these cases, the mixed mode estimator will ‘fall back’ to using frequency to estimate a probability of that edge being traversed. As implemented, the logic that makes this choice is in the estimator object itself and the graph is created with either a number of estimator as a weight. So all petersburg graphs now have the ability to accept pre-trained sklearn estimators as weights.
Conclusion
So there you have it. Finally petersburg has grown from a neat way to model some probability puzzles to a viable framework for understanding, modeling, and predicting complicated hierarchical events. I’m working on a more formalized, technical, in-depth document detailing the motivations for Petersburg, how it works, and how it can be used, but in the meantime, all of the code is up and working at https://github.com/wdm0006/petersburg so check it out.
The post Mixed-mode estimation in petersburg appeared first on Will’s Noise.